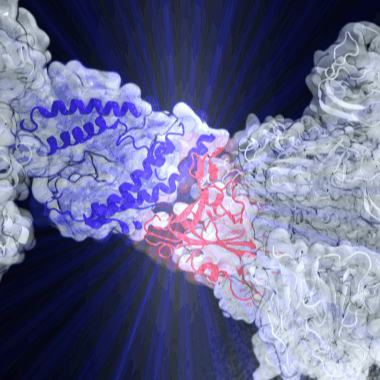
SARS-CoV-2 infections are initiated by attachment of the receptor-binding domain (RBD) on
the viral Spike protein to angiotensin-converting enzyme-2 (ACE2) on human host cells. We are combining experimental results with steered molecular dynamics simulations to investigate the mechanics of the RBD:ACE2 interaction. Our simulations can be used to describe how the virus have evolved to become more mechanically stable.
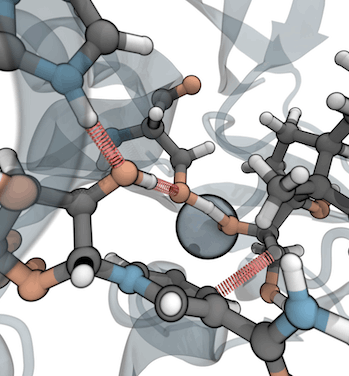
Combining hybrid QM/MM calculations with molecular biology experiments allows us to investigate the mechanisms
No Code Website Builder