What we do

Research
Software

About Us
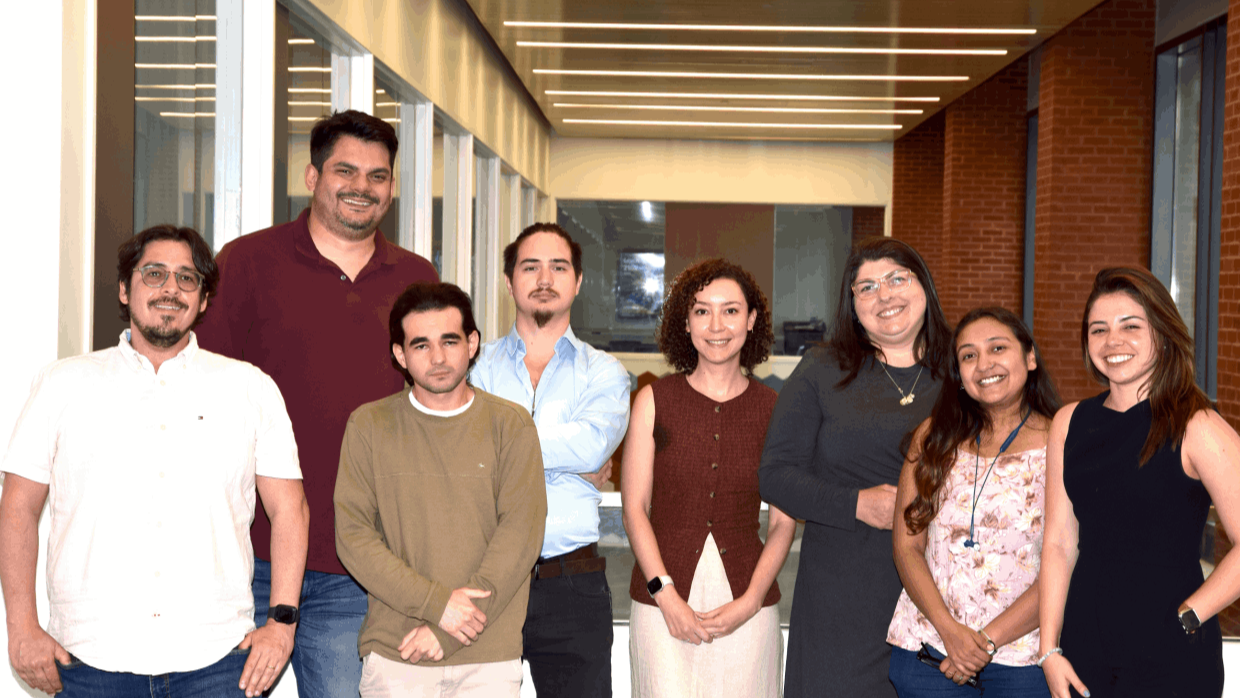

Building knowledge at the interface of Biology, Chemistry, and Physics.




AI Website Generator